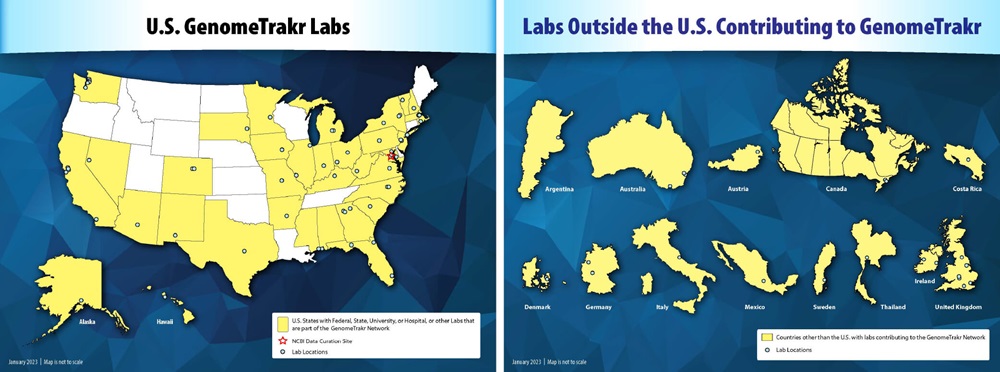
Domestic and international laboratories that participate in the GenomeTrakr network. (Source: FDA)
The GenomeTrakr Network helps quickly identify genetic sources of foodborne outbreaks. By connecting public health and university laboratories that utilize whole genome sequencing (WGS) for pathogen identification, GenomeTrakr allows these laboratories to share genomic and geographic data from foodborne outbreaks as they occur.
Contact the Food Safety team: [email protected]
GenomeTrakr labs perform whole genome sequencing bacteria isolates, as well as parasites and viruses. Food and environmental samples are sequenced and uploaded to the GenomeTrakr database by federal (e.g., FDA, USDA, CDC, NOAA), state, academic and international partners. Laboratories that participate in the GenomeTrakr Network use the data to identify foodborne outbreaks and contaminated products.

Domestic and international laboratories that participate in the GenomeTrakr network. (Source: FDA)
Galaxy Trakr is an open source, web-based platform for accessible, reproducible and transparent computational biomedical research. Laboratories that participate in the GenomeTrakr Network upload their whole genome sequencing data to the web-based platform to perform quality assessments of sequencing data, identify links between clinical isolates and positive food/environmental samples, including those at the National Center for Biotechnology Information sequence read archive and explore new methodologies such as metagenomics. It allows users without programming experience to easily specify parameters and run individual tools as well as larger workflows.
This tutorial walks the viewer through the four parts of the Galaxy Trakr interface: the tool list on the left, the viewing panel in the middle and the analysis and data history on the right.
This training video demonstrates how users can utilize Seqsero software to determine serotypes, or groups within a single species of microorganisms, such as bacteria or viruses, which share distinctive surface structures.